Diagnostic biomarker
A diagnostic biomarker refers to a biological measure that aids the diagnosis of a disease and may serve in determining disease progression and/or success of treatment. It may be a laboratory, radiological, genetic, anatomical, physiological or other finding that helps to differentiate one disease from others.
Defining a biomarker[edit | edit source]
In 2001, the World Health Organization (WHO), in coordination with the United Nations and the International Labor Organization, has defined a biomarker as “any substance, structure, or process that can be measured in the body or its products and influence or predict the incidence of outcome or disease.”[1] The Biomarkers Consortium, a major public-private biomedical research partnership, uses the 2001 National Institutes of Health (NIH) Biomarkers Definitions Working Group definition: "Biomarkers are characteristics that are objectively measured and evaluated as indicators of normal biological processes, pathogenic processes, or pharmacologic responses to therapeutic intervention."[2]
Presently, myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS) patients are diagnosed based on well-defined clinical criteria, for example, please see the Proposed Diagnostic Criteria Chart section of the Institute of Medicine report. Diagnosis also includes a process of exclusion of other causes of fatigue which can result in a delay of diagnosis.[3]
Potential diagnostic biomarkers for ME/CFS are being explored by many researchers in the field. Below is a list of some that have been considered. They are alphabetically ordered so as not to imply some may be more promising than others.
Blood tests[edit | edit source]
Activin B[edit | edit source]

A 2017 study that used the Canadian Consensus Criteria (CCC) concludes: "Elevated activin B levels together with normal activin A levels identified patients with the diagnostic symptoms of CFS/ME, thus providing a novel serum based test. The activins have multiple physiological roles and capture the diverse array of symptoms experienced by CFS/ME patients."[4]
The same group later used a weighted standing test in order to rate disease severity, and then compared subjects' health status to their levels of activin B. Activin B was once again found to separate people with ME/CFS from healthy controls.[5]
Buspirone challenge test[edit | edit source]
The buspirone challenge test measures how much prolactin is released into the bloodstream when a single dose of the drug buspirone (a 5-HT1A serotonin receptor agonist) is orally administered. This test has been shown to distinguish ME/CFS patients from healthy controls, as well as being able to distinguish ME/CFS patients from patients with depression.
CellTrend diagnostic test[edit | edit source]
CellTrend bases its test on the theory that a subset of 20-30% of all patients suffering from ME/CFS have developed elevated levels of three auto-antibodies, i.e., auto-antibodies against the b2-adrenergic receptor, auto-antibodies against the muscarinic cholinergic receptor 3 (M3) and auto-antibodies against the muscarinic cholinergic receptor 4 (M4).[6][7]
Obstacle for use: Only detects a subset 20-30%. Not widely accepted. Patients pay the test cost.
Cytokine expression[edit | edit source]
Several researchers are exploring if cytokine expression in ME/CFS is a unique enough signature to be used as a diagnostic marker.[8][9][10][11]
Cytokine expression changes in ME/CFS related to length of illness, with some cytokines levels increasing and some decreasing dependent on illness duration. Russell, et al, focused on a subset of three cytokines, IL-1α, 6 and 8, in plasma samples and concluded that: "Setting these 3 markers as a triple screen and adjusting their contribution according to illness duration [the] sub-groups produced ME/CFS classification accuracies of 75–88%."[8]
Obstacle for use: Since cytokine expression changes in ME/CFS related to progression of the illness, the validity of potentially useful markers may be obscured by such variation.[8]
In 2016, Landi, et al, "observed highly significant reductions in the concentration of circulating interleukin (IL)-16, IL-7, and Vascular Endothelial Growth Factor A (VEGF-A) in ME/CFS patients" but not in patients of other chronic illnesses with fatigue as a symptom. This three cytokine pattern is being studied further as a potential biomarker for ME/CFS.[12]
In 2018, Moneghetti, et al, compared the results of cytokine profiles 18 hours post exercise for ME/CFS patients vs healthy patients matched for cardiac structure and exercise capacity. They found that the most discriminatory cytokines detected post exercise in ME/CFS patients were CD40L, platelet activator inhibitor, interleukin 1-β, interferon-α and CXCL1. They concluded that cytokine profiling following exercise may help differentiate patients with ME/CFS from sedentary controls.[13]
Obstacle for use: These studies need to be duplicated with other patient cohorts to assure the results are specific enough for an accurate diagnosis of ME/CFS.
Dysfunction of TCA and urea cycles[edit | edit source]
A 2016 study in Japan, by Yamano, et al, looked at the differences in intermediate metabolite concentrations in the tricarboxylic acid (TCA) and urea cycles in CFS patients versus healthy controls: "CFS patients exhibited significant differences in intermediate metabolite concentrations in the tricarboxylic acid (TCA) and urea cycles. The combination of ornithine/citrulline and pyruvate/isocitrate ratios discriminated CFS patients from healthy controls, yielding area under the receiver operating characteristic curve values of 0.801 (95% confidential interval [CI]: 0.711–0.890, P < 0.0001) and 0.750 (95% CI: 0.584–0.916, P = 0.0069) for training (n = 93) and validation (n = 40) datasets, respectively. These findings provide compelling evidence that a clinical diagnostic tool could be developed for CFS based on the ratios of metabolites in plasma."[14]
Obstacle for use: The specialized laboratory equipment needed for this test is usually only available in facilities engaged in research. The findings in this published study need to be validated through replication by other studies.
EBV-encoded DNA polymerase and EBV-encoded dUTPase[edit | edit source]
In 2012, Dr Ronald Glaser and Dr A Martin Lerner, et al, discovered that the proteins, Epstein-Barr virus-encoded DNA polymerase and Epstein-Barr virus-encoded dUTPase, were present in the blood sera of an EBV subset of CFS patients, but was negative in sera of controls.[15]
Obstacle for use: These were preliminary findings that need to be are corroborated by studies with a larger number of EBV subset CFS patients.
Electrical Impedance[edit | edit source]
Experimentation is also being carried out at the End ME/CFS Project with the Open Medicine Foundation to test if a metabolic testing device using new nanofabricated technology that measures electrical impedance could be used to develop a simple diagnostic test for (ME/CFS). Blood from patients with ME/CFS causes a rapid and significant rise in electrical impedance when placed in this device, whereas blood from healthy people does not. The device is inexpensive and gives a real-time assay. This nanofabricated technology has the potential to be place in a hand-held device and disseminate to physician's offices for use as a test during medical visits.[16] A study on this was published in 2019, and described the nanoneedle designed for the test.[17]
Obstacle for use: The technology is still very new and needs further testing. If the technology works, development into an inexpensive, easily disseminated, handheld device is necessary.
Extracellular Vesicles[edit | edit source]
A 2018 study from Spain reported that extracellular vesicles (EVs) were significantly smaller as measured by protein cargo, size distribution and concentration, and the amount of EV-enriched fraction was significantly higher in the blood serum of CFS/ME patients versus healthy controls. Extracellular vesicles are secreted from both healthy cells and cells undergoing apoptosis (i.e., cell death) and play a role in intercellular communication.[18]
Obstacle for use: The study population consisted of 10 Spanish CFS/ME patients and 5 matched healthy controls. This study would need to be reproduced in larger populations to verify statistic significance.
Growth Differentiation Factor 15[edit | edit source]
GDF15 has been proposed as a biomarker for fatigue severity in ME/CFS patients.[19] Melvin et al (2019) used samples from the UK ME/CFS biobank to compare ME/CFS patients, and found fatigue severity positively correlated with severity of symptoms, and that higher GDF15 levels were found in ME/CFS patients, including mild/moderate patients, in comparison to either multiple sclerosis patients or healthy controls. The study used samples from 50 severe ME/CFS patients and 100 mild/moderate ME/CFS patients.[19] Results have not yet been replicated.
Immunosignatures[edit | edit source]
Immunosignaturing is a medical diagnostic test which uses arrays of random-sequence peptides (i.e., linked amino acids) to measure specific antibodies circulating in the blood. The Nevada Center for Biomedical Research in Reno, Nevada, US has identified a unique Immunosignature (IMS) comprised of a subset of 25 peptides that differentiates the blood serum from people with ME compared to controls with 92.9% specificity and 97.6% sensitivity. ME patients met both the International Consensus Criteria (CCC) and Fukuda criteria.[20]
Messenger RNA (mRNA)[edit | edit source]
In 2021, a Dutch study identified a signature of 23 genes capable of distinguishing between ME/CFS cases and healthy controls. They used publicly available mRNA expression data and and DNA methylation data from peripheral blood mononuclear cells (PBMCs) of 93 ME/CFS patients and 25 healthy controls. The authors believe that ten of the 23 genes could be interpreted in the context of the derailed immune system of ME/CFS. The ten proteins focused on in this study are MAPK4, ARRB1, GOLGA4, ABCE1, PHKA2, IL2RB, CCR4, HLA-DQA1, PRG4 and OGG1. The mRNA transcripts encoding these proteins were all downregulated in ME/CFS patients compared to healthy controls.[21]
Metabolomics[edit | edit source]
Naviaux, et al, at the University of California, San Diego School of Medicine reported in 2016 "that targeted, broad-spectrum metabolomics of plasma not only revealed a characteristic chemical signature but also revealed an unexpected underlying biology." The researchers found that CFS patients had an average of 40 metabolic abnormalities. From these, a set of 8 metabolites were identified in male patients and a set of 13 metabolites in females that performed well together as a diagnostic test. Both sets gave a 95% Confidence Interval. The eight metabolites selected for males were phosphatidyl choline PC(16:0/16:0), glucosylceramide GC(18:1/16:0), 1-P5C, FAD, pyroglutamic acid (also known as 5-oxoproline), 2-hydroxyisocaproic acid (HICA), l-serine, and lathosterol. The 13 metabolites selected for females were THC(18:1/24:0), phosphatidyl choline PC(16:0/16:0), hydroxyproline, ceramide(d18:1/22:2), lathosterol, adenosine, phosphatidylinositol PI(16:0/16:0), FAD, 2-octenoylcarnitine, phosphatidyl choline plasmalogen PC(22:6/P18:0), phosphatidyl choline PC(18:1/22:6), 1-P5C, and CDCA. There are several metabolites that appear on both the males and female lists.[22]
Obstacle for use: The specialized laboratory equipment needed for this test is usually only available in facilities engaged in research.
MicroRNA (miRNA)[edit | edit source]
microRNA (miRNA) are molecules involved in gene expression. (Note:MicroRNA are different from mRNA which stands for messenger RNA.)
A 2016 study by Petty, et al found that "four upregulated miRNA were suitable markers to resolve CFS/ME subjects from a matched control cohort."[23] A 2012 study by Brenu, et al found the expression of eight specific miRNAs "significantly decreased in NK cells of CFS/ME patients in comparison to the non-fatigued controls" and one specific miRNA was significantly downregulated in "both the NK and CD8(+)T cells in the CFS/ME sufferers."[24]
In 2015, Griffith University filed for a patent for a biological marker (Patent Publication number WO2016023077 A1) for the diagnosis and management of ME and CFS. Sonya Marshall-Gradisnik and Ekua Brenu are listed as the inventors. The patent application states: "The present invention resides broadly in the use of at least one miRNA as a biological marker for identifying or diagnosing a subject having CFS and/or ME."[25] In 2016, Griffith University's Professor Donald Staines and Professor Sonya Marshall-Gradisnik announced that they have been awarded a $4-million grant to be administered during the next five years that will enable them to continue research into developing a diagnostic test for ME/CFS.[26]
Almenar-Pérez et al (2020) analyzed microRNAs from both peripheral blood mononuclear cells and extracellular vesicles in 15 severely ill myalgic encephalomyelitis/
Although the science of using the detection of different miRNAs for disease confirmation is still in its infancy. There are no established baseline data for miRNAs among normal individuals, which would be necessary for using miRNA levels as biomarkers.[28] The findings in the studies published in March 2016 and January 2020 need to be validated through replication by other studies. The 2020 involved only severely ill women, and would need to be validated for use in children, men and those who are not severely ill. The Griffith University test has not been released for public use to date.
Mitochondrial energy production blockage[edit | edit source]
Studies by Myhill, Booth and McLaren-Howard[29][30][31] have shown the mitochondria of ME/CFS patients to have energy production defects. Their studies employed the ATP Profiles Test[32] (Acumen Laboratory, Tiverton, Devon, UK) to measure the functional efficiency of five metabolic processes involved in mitochondrial energy production (the test provides five numerical values indicating the functional efficiency of each energy metabolism process).
Using this ATP Profiles Test, the authors discovered that almost all the 200 or so ME/CFS patients in their cohorts had impaired mitochondrial energy production. However, out of the five energy metabolism processes measured, each patient had their own particular processes that were at fault (running at low efficiency), and their own particular processes which were working OK (running at normal efficiency).
Because these studies discovered ME/CFS patients can have energy metabolism defects in several of the energy metabolism processes measured, the authors combined the efficiency values for each of the 5 processes into one single efficiency figure, which they call the Mitochondrial Energy Score (MES). The MES is thus a single numerical value that gives the overall efficiency of a patient's mitochondrial energy production. The authors found there is a high degree of correlation between the Mitochondrial Energy Score and the degree of severity of ME/CFS (severity as measured on the Bell scale). Furthermore, the Mitochondrial Energy Score was able to successfully distinguish between ME/CFS patients and healthy controls in nearly all cases.
Nanoelectronics-blood-based diagnostic biomarker[edit | edit source]
A nanoelectronics-based biomarker for ME/CFS was found by a team lead by Ron Davis, at Stanford University, in 2019. This blood test uses electrical impedance testing and correctly identified 100% of patients with ME/CFS, diagnosed according to the Canadian Consensus Criteria.[17] Only 20 patients gave samples for this study, and 20 controls.[17] The blood test involves putting the white blood cell samples under physical stress, and relied on nanotechnology using a specially designed nanoneedle.[17]
Natural killer cell function[edit | edit source]
Numerous studies of CFS have found evidence of reduced natural killer cell (NK) function.[33][34][35][36][37] Some studies have showed NK function correlates with illness severity.[38] One study found increased differentiation in NK cells.[39]
Obstacles for use: Specialized lab equipment not available in average laboratories; blood specimen must be tested within 48 hours of draw and must remain at between 59°-98.6°F or 15°-37°C so special considerations are needed in transporting blood specimens to specialized labs. Reduced NK cell function can be seen with other immune-related conditions,[40] so this test would need to be paired with other clinical or laboratory findings to make a definitive diagnosis.
Phenylalanine measured via Raman microspectroscopy[edit | edit source]
A research team (Xu, et al, 2018) in the UK used single-cell Raman microspectroscopy (SCRM) on rho zero cells (ρ0 cells) that lack mitochondrial DNA (mtDNA) compared to peripheral blood mononuclear cells (PBMCs) from CFS patients and healthy controls. They found that Raman phenylalanine bands associated with CFS patient PBMCs were significantly higher and were increased in intensities compared to controls. The results suggest that the increase in cellular phenylalanine may relate to mitochondrial/energetic dysfunction in CFS and that phenylalanine can be used as a potential biomarker for diagnosis of CFS by SCRM. They achieved an accuracy rate of 98% correctly determining between CFS patients and healthy controls.[41]
Obstacles in use: Further testing needed for confirmation.
Plasma neuropeptide Y[edit | edit source]
Plasma levels of neuropeptide Y (NPY), a neurotransmitter in high quantity in the brain, are reported to be elevated in complex multi-symptom illnesses associated with [[immune system|immunologic] dysfunction. Fletcher, et al, did a study believed to be "the first in the CFS literature to report that plasma NPY is elevated [in ME/CFS] compared to healthy controls and to a fatigued comparison group, GWI [Gulf War Illness] patients. The significant correlations of NPY with stress, negative mood, general health, depression and cognitive function strongly suggest that this peptide be considered as a biomarker to distinguish subsets of CFS."[42]
Obstacles in use: NPY is elevated in other immune illnesses such as rheumatoid arthritis (RA) and systemic lupus erythematosus (SLE). NPY varies greatly between individuals and is interdependent on other cellular and molecular components and must be viewed with data from other biomarkers.[42]
Ratio of active:inactive PKR[edit | edit source]
In 2018, Eiren Sweetman discovered, during work for her doctorate thesis, that a changed ratio of active:inactive Protein Kinase R (PKR) in people with ME/CFS vs age-gender matched controls could potentially be used as a diagnostic biomarker. Protein Kinase R (PKR) is an innate antiviral immune response protein. Upon measuring an antibody against a non-phosphorylated PKR fragment and an antibody against a Thr-446 phosphorylated PKR peptide, an increased ratio of phosphorylated PKR to non-phosphorylated inactive PKR was detected ME/CFS patients.[43]
Obstacles in use: Further testing needed for confirmation; test would need to be adapted from a research setting to a clinical setting for wide availability.
Brain imaging[edit | edit source]
DFI and MRI scans[edit | edit source]
Using advanced brain imaging, Zeineh, et. al., found that there was right arcuate fasciculus (FA) abnormality in CFS patients. "Bilateral white matter atrophy is present in CFS...Right hemispheric increased FA may reflect degeneration of crossing fibers or strengthening of short-range fibers. Right anterior arcuate FA may serve as a biomarker for CFS."[44]
Obstacle for use: The specialized radiological equipment needed for this test is usually only available in facilities engaged in research. Radiologists would need to be trained to interpret the scans to identify the unique features present in ME/CFS brains.
ECG[edit | edit source]
Short QT interval[edit | edit source]
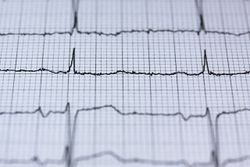
"Automated measurement of QTc [QT corrected] in clinical practise has potential utility as a diagnostic biomarker in CFS."[45]
The electrocardiographic QT interval is shorter in patients with ME/CFS than those of the general population.[45] The only exception is in those who have a rare genetic illness, Short QT syndrome. Differentiation is easy to determine because those with Short QT syndrome also have tall and peaked T waves, whereas the T waves are normal in people with ME. Modern computer-based ECG machines are programmed to correct the QT interval in relation to heart rate, because a number of medical conditions elicit a prolonged QT interval.
Obstacles for use: medical personnel education needed to recognize this finding on the EKG finding as indicative of ME/CFS; modern computer-based EKG machine needed
Cerebrospinal fluid[edit | edit source]
Orosomucoid[edit | edit source]
Orosomucoid (ORM) is a protein which has many biological activities including modulating immunity. In 2016, a study with Yang, et al, showed that the serum level of ORM in individuals with CFS was consistently elevated compared with that from healthy volunteers. The serum cortisol level in these same CFS patients was moderately decreased, indicating that "ORM increase is not a direct result of stress response."[46]
Orosomucoid 2 is one of the proteins Baraniuk, et al, found in a unique CFS – related proteome when studying pooled cerebrospinal fluid of CFS patients.[47]
Obstacles in use: Further testing needed for confirmation.
Proteome in cerebrospinal fluid[edit | edit source]
Analysis of the proteome (complement of proteins) in cerebrospinal fluid specimens by Baraniuk, et al, in 2005, showed one, consistent CFS-related proteome compared to a group with Gulf War Illness and to controls. They found the presence of 10 proteins in cerebrospinal fluid that were shared by patients with CFS, but were not detected in control samples."[47]
A similar study of proteome analysis of cerebrospinal fluid was done in 2011 by Steven Schutzer, et al. His group compared the pooled cerebrospinal fluid specimens of patients with CFS to patients with Neurologic Post Treatment Lyme disease (nPTLS) and to healthy controls. The cerebrospinal fluid proteome of the three groups were markedly unique for each group. Although nPTLS and CFS have similar clinical presentations, the researchers were able to distinguish the two syndromes from each other via data analysis.[48]
Obstacles in use: The specialized laboratory equipment needed for this test is usually only available in facilities engaged in research. The samples were pooled specimens. The presence of these proteins in one patient may be too small for average laboratory equipment to detect. Specific proteins to be used for a biomarker have not been identified. Cerebrospinal spinal fluid samples are obtained via spinal taps which are considered invasive and come with potential side effects.
Physiological tests [edit | edit source]
Hand grip strength[edit | edit source]
A 2018 UK ME/CFS biobank study led by Luis Nacul assessed over 200 patients with ME/CFS found that hand grip strength was both a possible biomarker for ME/CFS, and an indicator of disease severity. Hand grip strength results were compared with healthy controls and patients with Multiple Sclerosis.[49]
Jäkel et al. (2021) investigated repeat maximum handgrip strength (HGS) as a measure of muscle fatigability in 105 ME/CFS patients meeting the Canadian consensus criteria, compared to 18 with cancer-related fatigue (CRF) and healthy controls.[50]
Tilt table test results[edit | edit source]
Tilt table tests have shown abnormalities including an average 27% drop in blood flow to the brain in one study of bedbound with severe ME and orthostatic intolerance symptoms, using an angle of just 20 degrees.[51] Similar abnormalities in blood flow have been found in mild and moderate ME/CFS patients.[52][51]
Two-day cardiopulmonary exercise test[edit | edit source]
The two-day cardiopulmonary exercise test or 2-day CPET is an accepted, reliable test for post-exertional malaise (PEM), one of the cardinal symptoms that distinguishes between individuals with and without ME/CFS.[53]
Obstacles for use: The 2-day CPET "carries substantial risk for these patients as it may worsen their condition."[3] A single day CPET is a common test used as a cardiac stress test and therefore the equipment is widely available at medical centers, however, medical personnel would need to be educated on how to interpret a 2-day CPET test for ME/CFS diagnosis.
Saliva tests [edit | edit source]
Saliva proteins ACON and ATPB[edit | edit source]
Using whole saliva, Ciregia, et al, 2016, found two proteins, aconitate hydratase (ACON) and ATP synthase subunit beta (ATPB), were upregulated in CFS. When these two proteins were tested in combination, they gave a ROC curve with sensitivity of 93% and specificity of 70%, in a twin study where one twin was healthy and one had CFS. A second cohort of unrelated CFS patients and healthy controls, likewise proved ACON and ATPB as promising biomarkers.[54]
Saliva Fatigue Biomarker Index[edit | edit source]
A saliva test developed by (2022) combined a multiple values to produce an overall marker, which they named the Fatigue Biomarker Index (FBI).[55] Further studies are needed.[55]
Urine tests[edit | edit source]
Allantoin[edit | edit source]
A 2015 metabolic profiling study of female ME/CFS subjects by Armstrong, et al, showed an increase in allantoin in the urine collected as a first-void urine sample, i.e., collected upon arising. Allantoin has been considered a reliable indicator of exercise-induced oxidative stress in humans. Other studies have shown indications of oxidative stress in ME/CFS[56] "but this is the first to report of allantoin as a marker and the first to suggest a pathway linking ATP degradation and oxidative stress."[57]
Obstacle for use: The findings in this published study need to be validated through replication by other studies using a more heterogeneous ME/CFS population.
Combination of methods[edit | edit source]
Blood and muscle nerves (electrical)[edit | edit source]
A combination of M-wave, TBars, and CD26 values has been proposed as a possible diagnostic marker. A 2016 study in France by Fenouillet, et al, examined 36 ME/CFS cases and 11 healthy controls with regard to three biological variables: a) post-exercise M-wave - a measurement after electrical stimulation of muscle nerves which coordinates with muscle fatigue b) TBARS variations - a plasma marker which represents excessive oxidative stress response to exercise, and c) CD26-expression at rest - a protein potentially related to inflammation, which was found to decrease in CFS patients. Although the researchers set out to explore physiological and biological abnormalities that could be indicative of the etiology, "striking differences" in the results between patients and controls lead them to believe that this combination could be used to identify patients with ME/CFS and help to distinguish ME/CFS patients from fibromyalgia patients.[58]
Obstacle for use: The findings in this published study need to be validated through replication by other studies.
Blood and fecal matter[edit | edit source]
In 2016 gut microbiome study done by Giloteaux, Hanson, et. al., the gut bacterial microbiome of 48 ME/CFS patients and 39 healthy controls was examined by sequencing regions of 16S ribosomal ribonucleic acid (rRNA) genes that allow identification of the different types of bacteria present in the stool as well as inflammatory markers from serum. "Bacterial diversity was decreased in the ME/CFS specimens compared to controls, in particular, a reduction in the relative abundance and diversity of members belonging to the Firmicutes phylum...Using a machine learning approach trained on the data obtained from 16S rRNA and inflammatory markers, individuals were classified correctly as ME/CFS with a cross-validation accuracy of 82.93%."[59]
Others[edit | edit source]
Other biomarkers being researched:
- Dr. Kevin Conley and Dr. David Maughan, University of Washington, Seattle, Washington, US are studying the abnormal response of the muscle metabolites, NADP+/NADPH, as a molecular marker in ME/CFS patients before and after fatiguing exercise by using advanced, non-invasive magnetic resonance spectroscopy (MRS).[60]
- Dr. Isabel Barao hypothesizes that FcR (a protein found on the surface of certain cells) dysfunction in immune cells is a risk factor for the development of ME/CFS. The Bateman Horne Center, the National Cancer Institute (NCI) at NIH, and Roche Pharmaceuticals, are examining polymorphisms and mutations of FcgRs in NK cells of ME/CFS patients and their associations to antibody-dependent cell-mediated cytotoxicity (ADCC) capability and disease pathology.[61]
- the presence of salivary Human herpesvirus 6 (HHV6) and Human herpesvirus 7 (HHV7) biomarkers to distinguish physiological fatigue from pathological fatigue[62]
- visible and near-infrared (Vis-NIR) spectroscopy of patients' thumbs are being explored in Japan as a non-invasive test for CFS[63][64]
See also[edit | edit source]
Learn more[edit | edit source]
- 2012, Biomarkers for chronic fatigue.[65]
- 2014, Chronic Fatigue Syndrome: The Current Status and Future Potentials of Emerging Biomarkers. (Full text)[66]
- March 3, 2016, "Study proposes new name, definition and biomarkers for chronic fatigue syndrome" in ME Australia by Sasha Nimmo
- March 30, 2016, "Three new chronic fatigue syndrome and ME biomarker discoveries" in ME Australia by Sasha Nimmo
- 2017, "What, Exactly, is a Biomarker Anyway? And Why Don't We Have One for ME/CFS?" Solve ME/CFS Initiative explains Biomarkers in medicine for diagnosis and treatment.
- 2019, A nanoelectronics-blood-based diagnostic biomarker for myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS)[17] (Abstract)
References[edit | edit source]
- ↑ World Health Organization (2001), Biomarkers in Risk Assessment: Validity and Validation
- ↑ Biomarkers Definitions Working Group (2001), "Biomarkers and surrogate endpoints: Preferred definitions and conceptual framework", Clinical Pharmacology & Therapeutics, 200 (69): 89–95, doi:10.1067/mcp.2001.113989
- ↑ 3.0 3.1 Institute of Medicine (USA); Committee on the Diagnostic Criteria for Myalgic Encephalomyelitis/Chronic Fatigue Syndrome (February 10, 2015), Beyond Myalgic Encephalomyelitis/Chronic Fatigue Syndrome: Redefining an Illness
- ↑ Lidbury, Brett; Badia, Kita; Lewis, Donald; Hayward, Susan; Ludlow, Helen; Hedger, Mark; de Kretser, David (2017). "Activin B is a novel biomarker for chronic fatigue syndrome/myalgic encephalomyelitis (CFS/ME) diagnosis: a cross sectional study". Journal of Translational Medicine. doi:10.1186/s12967-017-1161-4.
- ↑ Richardson, Alice M.; Lewis, Don P.; Kita, Badia; Ludlow, Helen; Groome, Nigel P.; Hedger, Mark P.; de Kretser, David M.; Lidbury, Brett A. (April 12, 2018). "Weighting of orthostatic intolerance time measurements with standing difficulty score stratifies ME/CFS symptom severity and analyte detection". Journal of Translational Medicine. 16. doi:10.1186/s12967-018-1473-z. ISSN 1479-5876. PMC 5898049. PMID 29650052.
- ↑ "POTS - CFS/ME - CRPS". CellTrend Luckenwalde. Retrieved February 4, 2020.
- ↑ Loebel, M; Grabowski, P; Heidecke, H; Bauer, S; Hanitsch, LG; Wittke, K; Meisel, C; Reinke, P; Volk, H; Fluge, Ø; Mella, O; Scheibenbogen, C (2016), "Antibodies to β adrenergic and muscarinic cholinergic receptors in patients with Chronic Fatigue Syndrome", Brain, behavior, and immunity, 52: 32–39, doi:10.1016/j.bbi.2015.09.013
- ↑ 8.0 8.1 8.2 Russell, Lindsey; Broderick, Gordon; Taylor, Renee; Fernandes, Henrique; Harvey, Jeanna; Barnes, Zachary; Smylie, AnneLiese; Collado, Fanny; Balbin, Elizabeth; Katz, Ben; Klimas, Nancy; Fletcher, Mary Ann (2016). "Illness progression in chronic fatigue syndrome: a shifting immune baseline". BMC Immunology. 17 (3). doi:10.1186/s12865-016-0142-3.
- ↑ Peterson, D; Brenu, EW; Gottschalk, G; Ramos, SB; Nguyen, T; Staines, D; Marshall-Gradisnik, S (2015). "Cytokines in the Cerebrospinal Fluids of Patients with Chronic Fatigue Syndrome/Myalgic Encephalomyelitis". Mediators of Inflammation. 2015. doi:10.1155/2015/929720.
- ↑ Hornig, M; Gottschalk, G; Peterson, D; Knox, KK; Schultz, AF; Eddy, ML; Che, X; Lipkin, WI (2016), "Cytokine network analysis of cerebrospinal fluid in myalgic encephalomyelitis/chronic fatigue syndrome.", Molecular Psychiatry, 21 (2): 261–9, doi:10.1038/mp.2015.29
- ↑ Montoya, Jose G.; Holmes, Tyson H.; Anderson, Jill N.; Maecker, Holden T.; Rosenberg-Hasson, Yael; Valencia, Ian J.; Chu, Lily; Younger, Jarred W.; Tato, Cristina M.; Davis, Mark M. (2017), "Cytokine signature associated with disease severity in chronic fatigue syndrome patients", Proceedings of the National Academy of Sciences of the United States of America, 114 (34): E7150–E7158, doi:10.1073/pnas.1710519114
- ↑ Landi, Abdolamir; Broadhurst, David; Vernon, Suzanne D; Tyrrell, D. Lorne J.; Houghton, Michael (2016), "Reductions in circulating levels of IL-16, IL-7 and VEGF-A in myalgic encephalomyelitis/chronic fatigue syndrome", Cytokine, 78: 27–36, doi:10.1016/j.cyto.2015.11.018, PMID 26615570
- ↑ Moneghetti, Kegan J.; Skhiri, Mehdi; Contrepois, Kévin; Kobayashi, Yukari; Maecker, Holden T.; Davis, Mark M.; Snyder, Michael; Haddad, Francois; Montoya, Jose G. (2018). "Value of Circulating Cytokine Profiling During Submaximal Exercise Testing in Myalgic Encephalomyelitis/Chronic Fatigue Syndrome". Scientific Reports. 8. doi:10.1038/s41598-018-20941-w.
- ↑ Yamano, Emi; Sugimoto, Masahiro; Hirayama, Akiyoshi; Kume, Satoshi; Yamato, Masanori; Jin, Guanghua; Tajima, Seiki; Goda, Nobuhito; Iwai, Kazuhiro; Fukuda, Sanae; Yamaguti, Kouzi; Kuratsune, Hirohiko; Soga, Tomoyoshi; Watanabe, Yasuyoshi; Kataoka, Yosky (2016). "Index markers of chronic fatigue syndrome with dysfunction of TCA and urea cycles". Scientific Reports. 6 (34990). doi:10.1038/srep34990. PMID 27725700.
- ↑ Lerner, AM; Ariza, ME; Williams, M; Jason, L; Beqaj, S; Fitzgerald, JT; Lemeshow, S; Glaser, R (2012), "Antibody to Epstein-Barr Virus Deoxyuridine Triphosphate Nucleotidohydrolase and Deoxyribonucleotide Polymerase in a Chronic Fatigue Syndrome Subset", PLoS ONE, 7 (11): e47891, doi:10.1371/journal.pone.0047891
- ↑ An Update on ME/CFS Research with Dr. Ronald W. Davis, retrieved February 4, 2020
- ↑ 17.0 17.1 17.2 17.3 17.4 Davis, R. W.; Wilhelmy, J.; Nemat-Gorgani, M.; Kashi, A.; Esfandyarpour, R. (April 25, 2019). "A nanoelectronics-blood-based diagnostic biomarker for myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS)". Proceedings of the National Academy of Sciences. 116 (21): 10250–10257. doi:10.1073/pnas.1901274116. ISSN 0027-8424. PMID 31036648.
- ↑ Castro-Marrero, Jesús; Serrano-Pertierra, Esther; Oliveira-Rodríguez, Myriam; Zaragozá, Maria Cleofé; Martínez-Martínez, Alba; Blanco-López, María del Carmen; Alegre, José (2018), "Circulating extracellular vesicles as potential biomarkers in chronic fatigue syndrome/myalgic encephalomyelitis: an exploratory pilot study", Journal of Extracellular Vesicles, 7 (1), doi:10.1080/20013078.2018.1453730
- ↑ 19.0 19.1 Melvin, Audrey; Lacerda, Eliana; Dockrell, Hazel; O'Rahilly, Stephen; Nacul, Luis (November 27, 2019). "Circulating levels of GDF15 in patients with Myalgic Encephalomyelitis/Chronic Fatigue Syndrome". Journal of Translational Medicine. doi:10.17863/CAM.46401. ISSN 1479-5876. Retrieved December 1, 2019.
- ↑ Singh, Sahajpreet; Stafford, Phillip; Schlauch, Karen A.; Tillett, Richard R.; Gollery, Martin; Johnston, Stephen Albert; Khaiboullina, Svetlana F.; De Meirleir, Kenny L.; Rawat, Shanti; Mijatovic, Tatjana; Subramanian, Krishnamurthy; Palotás, András; Lombardi, Vincent C. (2016). "Humoral Immunity Profiling of Subjects with Myalgic Encephalomyelitis Using a Random Peptide Microarray Differentiates Cases from Controls with High Specificity and Sensitivity". Molecular Neurobiology. doi:10.1007/s12035-016-0334-0.
- ↑ Metselaar, Paula I.; Mendoza-Maldonado, Lucero; Li Yim, Andrew Yung Fong; Abarkan, Ilias; Henneman, Peter; te Velde, Anje A.; Schönhuth, Alexander; Bosch, Jos A.; Kraneveld, Aletta D. (February 2021). "Recursive ensemble feature selection provides a robust mRNA expression signature for myalgic encephalomyelitis/chronic fatigue syndrome". Scientific Reports. 11 (1): 4541. doi:10.1038/s41598-021-83660-9. ISSN 2045-2322. PMC 7907358. PMID 33633136.
- ↑ Petty, Robert D.; McCarthy, Neil E.; Le Dieu, Rifca; Kerr, Jonathan R. (March 11, 2016). "MicroRNAs hsa-miR-99b, hsa-miR-330, hsa-miR-126 and hsa-miR-30c: Potential Diagnostic Biomarkers in Natural Killer (NK) Cells of Patients with Chronic Fatigue Syndrome (CFS)/ Myalgic Encephalomyelitis (ME)". PLOS ONE. doi:10.1371/journal.pone.0150904. PMID 26967895.
- ↑ Brenu, Ekua W; Ashton, KJ; van Driel, M; Staines, Donald; Peterson, Daniel; Atkinson, GM; Marshall-Gradisnik, Sonya (2012). "Cytotoxic lymphocyte microRNAs as prospective biomarkers for Chronic Fatigue Syndrome/Myalgic Encephalomyelitis". Journal of Affective Disorders. 141 (2–3): 261–9. doi:10.1016/j.jad.2012.03.037.
- ↑ Biological markers, retrieved February 4, 2020
- ↑ "Coast team zeroing in on CFS test". goldcoastbulletin.com.au. December 1, 2016. Retrieved February 4, 2020.
- ↑ Almenar-Pérez, Eloy; Sarría, Leonor; Nathanson, Lubov; Oltra, Elisa (February 7, 2020). "Assessing diagnostic value of microRNAs from peripheral blood mononuclear cells and extracellular vesicles in Myalgic Encephalomyelitis/Chronic Fatigue Syndrome". Scientific Reports. 10 (1): 1–15. doi:10.1038/s41598-020-58506-5. ISSN 2045-2322. Retrieved February 12, 2020.
- ↑ Tonge, D.P.; Gant, T.W. (2016), "What is normal? Next generation sequencing-driven analysis of the human circulating miRNAOme", BMC Molecular Biology, 17 (4), doi:10.1186/s12867-016-0057-9
- ↑ Myhill, Sarah; Booth, Norman E.; McLaren-Howard, John (January 15, 2009). "Chronic fatigue syndrome and mitochondrial dysfunction". International Journal of Clinical and Experimental Medicine. 2 (1): 1–16. ISSN 1940-5901. PMC 2680051. PMID 19436827.
- ↑ Booth, Norman E; Myhill, Sarah; McLaren-Howard, John (June 15, 2012). "Mitochondrial dysfunction and the pathophysiology of Myalgic Encephalomyelitis/Chronic Fatigue Syndrome (ME/CFS)". International Journal of Clinical and Experimental Medicine. 5 (3): 208–220. ISSN 1940-5901. PMC 3403556. PMID 22837795.
- ↑ Myhill, Sarah; Booth, Norman E; McLaren-Howard, John (November 20, 2012). "Targeting mitochondrial dysfunction in the treatment of Myalgic Encephalomyelitis/Chronic Fatigue Syndrome (ME/CFS) - a clinical audit". International Journal of Clinical and Experimental Medicine. 6 (1): 1–15. ISSN 1940-5901. PMC 3515971. PMID 23236553.
- ↑ "Mitochondrial Function Profile - DoctorMyhill". drmyhill.co.uk. Retrieved February 4, 2020.
- ↑ Barker, Edward; Fujimura, Sue F.; Fadem, Mitchell B; Landay, Alan L.; Levy, Jay A. (1994). "Immunologic Abnormalities Associated with Chronic Fatigue Syndrome". Clin Infect Dis. 18: S136–S141. doi:10.1093/clinids/18.Supplement_1.S136.
- ↑ Whiteside, TL; Friberg, D (1998), "Natural killer cells and natural killer cell activity in chronic fatigue syndrome.", Am J Med, 105 (3A): 27S–34S, PMID 9790479
- ↑ Brenu, EW; Huth, TK; Hardcastle, SL; Fuller, K; Kaur, M; Johnston, S; Ramos, Sandra; Staines, Donald; Marshall-Gradisnik, Sonya (2014). "Role of adaptive and innate immune cells in chronic fatigue syndrome/myalgic encephalomyelitis". Int Immunol. 26 (4): 233–42. doi:10.1093/intimm/dxt068. PMID 24343819.
- ↑ Fletcher, Mary Ann; Maher, Kevin J; Klimas, Nancy (April 2002), "Natural killer cell function in chronic fatigue syndrome", Clinical and Applied Immunology Reviews, 2 (2): 129–139, doi:10.1016/S1529-1049(01)00047-2
- ↑ Brenu, Ekua W; van Driel, Mieke L; Staines, Donald R; Ashton, Kevin J; Hardcastle, Sharni L; Keane, James; Tajouri, Lotti; Peterson, Daniel; Ramos, Sandra B; Marshall-Gradisnik, Sonya M (2012). "Longitudinal investigation of natural killer cells and cytokines in chronic fatigue syndrome/myalgic encephalomyelitis". Journal of Translational Medicine. 10: 88. doi:10.1186/1479-5876-10-88.
- ↑ Ojo-Amaize, Emmanuel A.; Conley, Edward J.; Peter, James B. (1994). "Decreased Natural Killer Cell Activity Is Associated with Severity of Chronic Fatigue Immune Dysfunction Syndrome". Clin Infect Dis. 18 (Supplement 1): S157–S159. doi:10.1093/clinids/18.Supplement_1.S157.
- ↑ Huth, TK; Brenu, EW; Ramos, S; Nguyen, T; Broadley, S; Staines, D; Marshall-Gradisnik, S (January 2016), "Pilot Study of Natural Killer Cells in Chronic Fatigue Syndrome/Myalgic Encephalomyelitis and Multiple Sclerosis", Scand J Immunol, 83 (1): 44–51, doi:10.1111/sji.12388, PMID 26381393
- ↑ Orange, J. S. (2013). Natural killer cell deficiency. The Journal of Allergy and Clinical Immunology, 132(3), 515–526. http://doi.org/10.1016/j.jaci.2013.07.020
- ↑ Xu, Jiabao; Potter, Michelle; Tomas, Cara; Elson, Jo; Morten, Karl; Poulton, Joanna; Wang, Ning; Jin, Hanqing; Hou, Zhaoxu (2018). "A new approach to find biomarkers in chronic fatigue syndrome/myalgic encephalomyelitis (CFS/ME) by single-cell Raman micro-spectroscopy". The Analyst. doi:10.1039/C8AN01437J. ISSN 0003-2654.
- ↑ 42.0 42.1 Fletcher, Mary Ann; Rosenthal, Martin; Antoni, Michael; Ironson, Gail; Zeng, Xiao R; Barnes, Zachary; Harvey, Jeanna M; Hurwitz, Barry; Levis, Silvina; Broderick, Gordon; Klimas, Nancy G (2010). "Plasma neuropeptide Y: a biomarker for symptom severity in chronic fatigue syndrome". Behavioral and Brain Functions. 6 (76). doi:10.1186/1744-9081-6-76.
- ↑ Sweetman, E. C. (2018). Comprehensive molecular analysis of different classes of molecules in a Myalgic Encephalomyelitis/Chronic Fatigue Syndrome pilot study group, and investigation of RNA-activated Protein Kinase R (PKR) as a diagnostic biomarker (Thesis, Doctor of Philosophy). University of Otago. Retrieved from http://hdl.handle.net/10523/7834
- ↑ Zeineh, Michael M.; Kang, James; Atlas, Scott W.; Raman, Mira M.; Reiss, Allan L.; Norris, Jane L.; Valencia, Ian; Montoya, Jose G. (February 2015), "Right Arcuate Fasciculus Abnormality in Chronic Fatigue Syndrome" (PDF), Radiology, 274 (2): 517–526, doi:10.1148/radiol.14141079
- ↑ 45.0 45.1 Scott, Ashley; Norton, Michael; Mabillard, Holly; Newton, Julia (2012), "Shortened QTc interval in chronic fatigue syndrome" (PDF), Bulletin of IACFS/ME, 19 (3/4): 202–211
- ↑ Yang, Sun; Zhen-Xian, Zhang; Liu, Xia (2016), "Orosomucoid (ORM) as a Potential Biomarker for the Diagnosis of Chronic Fatigue Syndrome (CFS)", CNS Neuroscience & Therapeutics, 22 (3): 251–252, doi:10.1111/cns.12522
- ↑ 47.0 47.1 Baraniuk, J.B.; Casado, B.; Maibach, H.; Clauw, D.J.; Pannell, L.K.; Hess, S.A. (2005), "Chronic fatigue syndrome – related proteome in human cerebrospinal fluid", BMC Neurology, 5 (22), doi:10.1186/1471-2377-5-22
- ↑ Schutzer, Steven E; Angel, Thomas E.; Liu, Tao; Schepmoes, Athena A.; Clauss, Therese R.; Adkins, Joshua N.; Camp, David G.; Holland, Bart K.; Bergquist, Jonas; Coyle, Patricia K.; Smith, Richard D.; Fallon, Brian A.; Natelson, Benjamin (2011). "Distinct Cerebrospinal Fluid Proteomes Differentiate Post-Treatment Lyme Disease from Chronic Fatigue Syndrome". PLoS ONE. 6 (2). doi:10.1371/journal.pone.0017287.
- ↑ Nacul, Luis Carlos; Mudie, Kathleen; Kingdon, Caroline; Clark, Taane G.; Lacerda, Eliana Mattos (2018). "Hand grip strength as a clinical biomarker for ME/CFS and disease severity". Frontiers in Neurology. 9. doi:10.3389/fneur.2018.00992. ISSN 1664-2295.
- ↑ Jäkel, Bianka; Kedor, Claudia; Grabowski, Patricia; Wittke, Kirsten; Thiel, Silvia; Scherbakov, Nadja; Doehner, Wolfram; Scheibenbogen, Carmen; Freitag, Helma (April 19, 2021). "Hand grip strength and fatigability: correlation with clinical parameters and diagnostic suitability in ME/CFS". Journal of Translational Medicine. 19 (1): 159. doi:10.1186/s12967-021-02774-w. ISSN 1479-5876. PMC 8056497. PMID 33874961.
- ↑ 51.0 51.1 van Campen, C. (Linda) M. C.; Rowe, Peter C.; Visser, Frans C. (June 2020). "Cerebral Blood Flow Is Reduced in Severe Myalgic Encephalomyelitis/Chronic Fatigue Syndrome Patients During Mild Orthostatic Stress Testing: An Exploratory Study at 20 Degrees of Head-Up Tilt Testing". Healthcare. 8 (2): 169. doi:10.3390/healthcare8020169.
- ↑ van Campen, C. (Linda) M.C.; Verheugt, Freek W.A.; Rowe, Peter C.; Visser, Frans C. (February 8, 2020). "Cerebral blood flow is reduced in ME/CFS during head-up tilt testing even in the absence of hypotension or tachycardia: A quantitative, controlled study using Doppler echography". Clinical Neurophysiology Practice. 5: 50–58. doi:10.1016/j.cnp.2020.01.003. PMC 7044650. PMID 32140630.
- ↑ Davenport, TE; Stevens, SR; Baroni, K; Van Ness, M; Snell, CR (2011), "Diagnostic accuracy of symptoms characterising chronic fatigue syndrome.", Disability and Rehabilitation, 33 (19–20): 1768–75, doi:10.3109/09638288.2010.546936
- ↑ Ciregia, F; Kollipara, L; Giusti1, L; Zahedi, RP; Giacomelli, C; Mazzoni, M R; Giannaccini, G; Scarpellini, P; Urbani, A; Sickmann, A; Lucacchini, A; Bazzichi, L (2016), "Bottom-up proteomics suggests an association between differential expression of mitochondrial proteins and chronic fatigue syndrome", Translational Psychiatry, 6 (9): e904, doi:10.1038/tp.2016.184
- ↑ 55.0 55.1 Jason, Leonard A.; Kalns, John; Richarte, Alicia; Katz, Ben Z.; Torres, Chelsea (October 2, 2021). "Saliva fatigue biomarker index as a marker for severe myalgic encephalomyelitis/chronic fatigue syndrome in a community based sample". Fatigue: Biomedicine, Health & Behavior. 9 (4): 189–195. doi:10.1080/21641846.2021.1994222. ISSN 2164-1846. PMC 8855987. PMID 35186443.
- ↑ Morris, Gerwyn; Maes, Michael (2014), "Oxidative and Nitrosative Stress and Immune-Inflammatory Pathways in Patients with Myalgic Encephalomyelitis (ME)/Chronic Fatigue Syndrome (CFS)", Current Neuropharmacology, 12 (2): 168–185, doi:10.2174/1570159X11666131120224653
- ↑ Armstrong, Christopher W.; McGregor, Neil R.; Lewis, Donald P.; Butt, Henry L.; Gooley, Paul R. (2015). "Metabolic profiling reveals anomalous energy metabolism and oxidative stress pathways in chronic fatigue syndrome patients". Metabolomics. 11 (6): 1626–1639. doi:10.1007/s11306-015-0816-5.
- ↑ Fenouillet, E.; Vigouroux, A.; Steinberg, J.G.; Chagvardieff, A.; Retornaz, F.; Guieu, R.; Jammes, Y. (2016), "Association of biomarkers with health-related quality of life and history of stressors in myalgic encephalomyelitis/chronic fatigue syndrome patients", Journal of Translational Medicine, 14 (1): 251, doi:10.1186/s12967-016-1010-x, PMID 27580693
- ↑ Giloteaux, L.; Goodrich, J.K.; Walters, W.A.; Levine, S.M.; Ley, R.; Hanson, M.R. (2016), "Reduced diversity and altered composition of the gut microbiome in individuals with myalgic encephalomyelitis/chronic fatigue syndrome.", Microbiome, 4 (1): 30, doi:10.1186/s40168-016-0171-4, PMID 27338587
- ↑ "Non-Invasive Diagnostic Test Project Seeks to Detect ME/CFS". Solve ME/CFS Initiative. January 13, 2017. Retrieved February 4, 2020.
- ↑ "Uncovering Biomarkers - Fcg Receptors in ME/CFS". Bateman Horne Center. April 15, 2016. Retrieved February 4, 2020.
- ↑ Aoki, R; Kobayashi, N; Suzuki, G; Kuratsune, H; Shimada, K; Oka, N; Takahashi, M; Yamadera, W; Iwashita, M; Tokuno, S; Nibuya, M; Tanichi, M; Mukai, Y; Mitani, K; Kondo, K; Ito, H; Nakayama, K (2016), "Human herpesvirus 6 and 7 are biomarkers for fatigue, which distinguish between physiological fatigue and pathological fatigue", Biochemical and Biophysical Research Communications, 478 (1): 424–30, doi:10.1016/j.bbrc.2016.07.010, PMID 27396623
- ↑ Sakudo, A. (2016). "Potential use of visible and near-infrared spectroscopy for the analysis and diagnosis of chronic fatigue syndrome (Review)". Molecular Medicine Reports. 14 (3): 1875–1879. doi:10.3892/mmr.2016.5476.
- ↑ Sakudo, A.; Kuratsune, H.; Kato, YH.; Ikuta, K. (2012). "Visible and near-infrared spectra collected from the thumbs of patients with chronic fatigue syndrome for diagnosis". Clinica Chimica Acta. 413 (19–20): 1629–32. doi:10.1016/j.cca.2012.05.004. PMID 22583968.
- ↑ Klimas, NG; Broderick, G; Fletcher, MA (2012), "Biomarkers for chronic fatigue.", Brain Behav Immun, 26 (8): 1202–10, doi:10.1016/j.bbi.2012.06.006, PMID 22732129
- ↑ Fischer, David B.; William, Arsani H.; Strauss, Adam C.; Unger, Elizabeth R.; Jason, Leonard; Marshall, Jr, Gailen D.; Dimitrakoff, Jordan D. (2014), "Chronic Fatigue Syndrome: The Current Status and Future Potentials of Emerging Biomarkers", Fatigue: Biomedicine, Health & Behavior, 2 (2): 93–109, doi:10.1080/21641846.2014.906066

