Genetics of chronic fatigue syndrome
The genes linked to myalgic encephalomyelitis/chronic fatigue syndrome is an area of research as ME/CFS has been observed in families.[1] It is unknown if there is a genetic link or common environmental exposure (infectious or toxic). Studies of twins show higher rates of ME/CFS in identical than fraternal twins. The Centers for Disease Control and Prevention (CDC) notes that specific genetic associations have not been established.[1]
The IDO Metabolic Trap Hypothesis for the Etiology of ME/CFS[edit | edit source]
A new hypothesis, the indolamine-2,3-dioxygenase (IDO) metabolic trap, was developed and formulated as a mathematical model. The historical occurrence of ME/CFS outbreaks is a singular feature of the disease and implies that any predisposing genetic mutation must be common. A database search for common damaging mutations in human enzymes produces 208 hits, including IDO2 with four such mutations. Non-functional IDO2, combined with well-established substrate inhibition of IDO1 and kinetic asymmetry of the large neutral amino acid transporter, LAT1, yielded a mathematical model of tryptophan metabolism that displays both physiological and pathological steady-states. Escape from the pathological one requires an exogenous perturbation. This model also identifies a critical point in cytosolic tryptophan abundance beyond which descent into the pathological steady-state is inevitable.[2]
ME/CFS Gene Study[edit | edit source]
The ME/CFS Gene Study is still collecting data, but the initial pilot study by Perez et al. (2019) found 10 relatively common genes or gene variants were significantly more common in people with ME/CFS.[3] These were CYP2D6, PRRT4, PRSS56, C14orf37, ANKDD1B, GPBAR1, LHB, ADAMTS19, VARS2, and CPLX2.[3]
In 2021 the pilot was reviewed and found to have had serious methodological flaws that rendered its findings incorrect and conclusions spurious.[4] It is not clear how a study with such obvious and serious problems might even have passed peer review.
Utah Population Database study[edit | edit source]
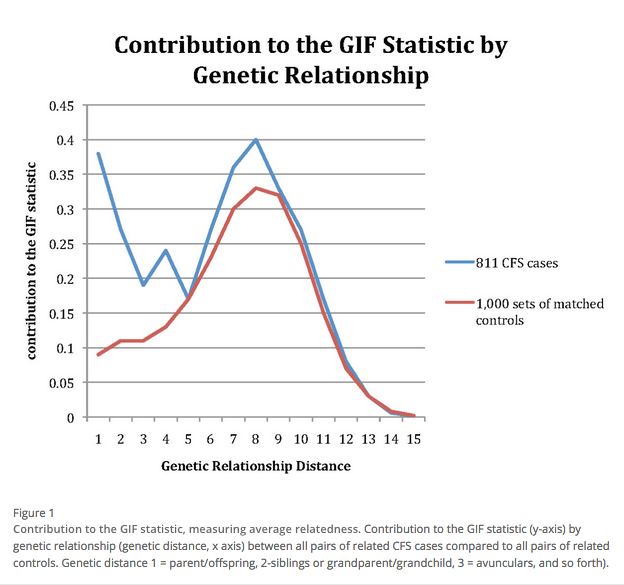
A 2011 study by Albright et al showed evidence of a heritable contribution to chronic fatigue syndrome (CFS).[5] Using the extensive records of the Utah Population Database (UPDB), the study "shows clear evidence of significant excess familial clustering and significantly elevated risks for CFS among first, second, and third degree relatives of CFS cases. The results strongly support a genetic contribution to predisposition to CFS as it is currently defined and diagnosed by clinicians in Utah." Increased outbreak rates in first degree relatives are not automatically assumed to be genetic because the first degree relatives often share the same lifestyle and environment. However, a significantly increased incidence (95% confidence interval) in second and third degree relatives strongly indicated a genetic contribution to CFS, given the much lower likelihood of these relatives sharing common risks and environments.[6]
Family history and twin studies[edit | edit source]
A 2001 study in the UK showed "there were significantly higher rates of CFS in the relatives of CFS cases compared with the relatives of control subjects."[7] Three twin studies (one in Australia, one in Washington, US, both in 2001, and one in the UK in 2007) showed that the correlations for prolonged and chronic fatigue were significantly higher in monozygotic than dizygotic twins for each definition of chronic fatigue syndrome.[8][9][10]
Haplogroups[edit | edit source]
One study showed that patients with mitochondrial DNA from certain haplogroups correlated with variations in gastrointestinal, neurological, and inflammatory symptoms.[11]
HLA alleles[edit | edit source]
Human leukocyte antigen genes associations were investigated by Lande et al. (2020) because these gene variants are considered hallmarks for autoimmune disease; two HLA associations were found to be more common in ME/CFS patients, but the majority of ME/CFS patients did not have these.[12]
Other genes[edit | edit source]
Glutamate receptor, ionotropic, kinase 2 (GRIK2) and neuronal PAS domain protein 2 (NPAS2) were also investigated in a CDC by Smith et al. (2011).[13]
A review by Dibble, McGrath and Ponting (2020) in preparation for the DecodeME study stated that research on the UK Biobank patient data showed associations for SLC25A15 (rs7337312), P4HA1 (rs150954845), EBF3 (rs148723539) and COX7B2 (rs564809936).[14]
The largest genetic study of ME/CFS patients was published by Hajdarevic et al. in 2022; and analysed data from over 2,500 patients from the UK Biobank, Norway and Denmark, reported they "did not find any ME/CFS risk loci displaying genome-wide significance". The study did find that the Tubulin Polymerization Promoting Protein (TPPP) region may be implicated in ME/CFS.[15][16] TPPP is an antibody expressed in the brain. Other genes identified LINC00333, RIN3, IGFBP1/IGFBP3, IZUMO1/FUT1 and ZBTB46 but PTPN22 and CTLA4 genes reported in previous research did not show significance.[15] Hajdarevic and colleagues found more significant associations within the Norwegian cohort, who (like the Danish cohort) were diagnosed using the strict Canadian Consensus Criteria, whereas the majority of data came from UK patient self-reported diagnosis. They commented that statistical significance would need approximately 10 times more patient data, which the DecodeME project was hoped to provide.[15]
Notable studies[edit | edit source]
- 2001, A twin study of the etiology of prolonged fatigue and immune activation[9] (Abstract)
- 2001, A twin study of chronic fatigue[8] (Abstract)
- 2006, Combinations of single nucleotide polymorphisms in neuroendocrine effector and receptor genes[17] (Full text)
Three genes were found to be common in a group of people with Chronic Fatigue Syndrome compared to the general population; TPH2 - neuronal tryptophan hydroxylase, COMT - catechol-O-methyltransferase, and NR3C1 - nuclear receptor subfamily 3, group C, member 1 glucocorticoid receptor, together these three have an accuracy of 76%.[17]
- 2007, Twin analyses of fatigue[10]
- 2009, A gene signature for post-Infectious chronic fatigue syndrome[18] (Full Text)
- 2009, Moderate Exercise Increases Expression for Sensory, Adrenergic, and Immune Genes in Chronic Fatigue Syndrome Patients But Not in Normal Subjects[19] (Full Text)
- 2011, Evidence for a heritable predisposition to Chronic Fatigue Syndrome[20] (Abstract)
- 2011, Gene expression alterations at baseline and following moderate exercise in patients with Chronic Fatigue Syndrome and Fibromyalgia Syndrome[21] (Full text)
- 2011, Convergent Genomic Studies Identify Association of GRIK2 and NPAS2 with Chronic Fatigue Syndrome[13] (Full text)
- 2016, Genome-wide association analysis identifies genetic variations in subjects with myalgic encephalomyelitis/chronic fatigue syndrome[22] (Full Text)
- 2016, Mitochondrial DNA variants correlate with symptoms in myalgic encephalomyelitis/chronic fatigue syndrome [11] (Abstract)
- 2018, Identification of Myalgic Encephalomyelitis/Chronic Fatigue Syndrome-associated DNA methylation patterns[23] (Abstract)
- 2018, Genome-epigenome interactions associated with Myalgic Encephalomyelitis/Chronic Fatigue Syndrome.[24] (Abstract)
- 2019, Associations between clinical symptoms, plasma norepinephrine and deregulated immune gene networks in subgroups of adolescent with Chronic Fatigue Syndrome.[25] (Abstract)
- 2019, Genetic Predisposition for Immune System, Hormone, and Metabolic Dysfunction in Myalgic Encephalomyelitis/Chronic Fatigue Syndrome: A Pilot Study[3] (Full text)
- 2019, The IDO Metabolic Trap Hypothesis for the Etiology of ME/CFS[2] - (Full text)
- 2020, Human Leukocyte Antigen alleles associated with Myalgic Encephalomyelitis/Chronic Fatigue Syndrome (ME/CFS)[12] - (Full text)
- 2020, Unravelling myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS): Gender‐specific changes in the microRNA expression profiling in ME/CFS[26] - (Full text)
- 2020, Genetic risk factors of ME/CFS: a critical review[14] - (Full text)
- 2021, Bioenergetic and Proteomic Profiling of Immune Cells in Myalgic Encephalomyelitis/Chronic Fatigue Syndrome Patients: An Exploratory Study[27] - (Full text)
- 2021, Re-analysis of Genetic Risks for Chronic Fatigue Syndrome From 23andMe Data Finds Few Remain[28]
- 2022, Genetic association study in myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS) identifies several potential risk loci[15] - (Full text)
Media Coverage[edit | edit source]
- 2016, Genome-wide associations
- 2016, New study found SNPs for some symptoms (CFS Remission, January 23)
See also[edit | edit source]
Learn more[edit | edit source]
- Mitochondrial DNA and ME/CFS - A Guide to the Hanson Lab's - 2016 JTM Publication
- The IDO Metabolic Trap Hypothesis For ME/CFS - Jul 30, 2019 Solve M.E.
- Dr. Ron Davis Gives an Update on ME/CFS Research- Sep 26, 2019 Open Medicine Foundation (Gene Mutation Found)
References[edit | edit source]
- ↑ 1.0 1.1 "Etiology and Pathophysiology | Presentation and Clinical Course | Healthcare Providers | Myalgic Encephalomyelitis/Chronic Fatigue Syndrome". Centers for Disease Control and Prevention. November 8, 2018. Retrieved February 8, 2019.
- ↑ 2.0 2.1 Kashi, Alex A.; Davis, Ronald W.; Phair, RobertD. (September 2019). "The IDO Metabolic Trap Hypothesis for the Etiology of ME/CFS". Diagnostics. 9 (3): 82. doi:10.3390/diagnostics9030082.
- ↑ 3.0 3.1 3.2 Nathanson, Lubov; Craddock, Travis J.A.; Klimas, Nancy G.; Gemayel, Kristina; Del Alamo, Ana; Hilton, Kelly; Jaundoo, Rajeev; Perez, Melanie (2019). "Genetic Predisposition for Immune System, Hormone, and Metabolic Dysfunction in Myalgic Encephalomyelitis/Chronic Fatigue Syndrome: A Pilot Study". Frontiers in Pediatrics. 7: 206. doi:10.3389/fped.2019.00206. ISSN 2296-2360.
- ↑ Bedford, Felice L.; Greshake Tzovaras, Bastian (March 18, 2021). "Re-analysis of Genetic Risks for Chronic Fatigue Syndrome From 23andMe Data Finds Few Remain". Frontiers in Pediatrics. 9: 590040. doi:10.3389/fped.2021.590040. ISSN 2296-2360. PMC 8012483. PMID 33816394.
- ↑ Albright, Frederick; Light, Kathleen; Light, Alan; Bateman, Lucinda; Cannon-Albright, Lisa A. (May 27, 2011). "Evidence for a heritable predisposition to Chronic Fatigue Syndrome". BMC neurology. 11: 62. doi:10.1186/1471-2377-11-62. ISSN 1471-2377. PMC 3128000. PMID 21619629.
- ↑ Albright, Frederick; Light, Kathleen; Light, Alan; Bateman, Lucinda; Cannon-Albright, Lisa A. (May 27, 2011). "Evidence for a heritable predisposition to Chronic Fatigue Syndrome". BMC neurology. 11: 62. doi:10.1186/1471-2377-11-62. ISSN 1471-2377. PMC 3128000. PMID 21619629.
- ↑ Walsh, C.M.; Zainal, N. Z.; Middleton, S.J.; Paykel, E. S. (September 2001). "A family history study of chronic fatigue syndrome". Psychiatric Genetics. 11 (3): 123–128. doi:10.1097/00041444-200109000-00003. ISSN 0955-8829.
- ↑ 8.0 8.1 Buchwald, D.; Herrell, R.; Ashton, S.; Belcourt, M.; Schmaling, K.; Sullivan, P.; Neale, M.; Goldberg, J. (2001). "A twin study of chronic fatigue". Psychosomatic Medicine. 63 (6): 936-943. PMID 11719632.
- ↑ 9.0 9.1 Hickie, IB; Bansal, AS; Kirk, KM; Lloyd, AR; Martin, NG (2001). "A twin study of the etiology of prolonged fatigue and immune activation". Twin Research. 4 (2): 94-102. doi:10.1375/1369052012209. PMID 11665341.
- ↑ 10.0 10.1 Schur, Ellen; Afari, Niloofar; Goldberg, Jack; Buchwald, Dedra; Sullivan, Patrick F. (2007). "Twin analyses of fatigue". Twin Research and Human Genetics. 10 (5): 729-733. doi:10.1375/twin.10.5.729.
- ↑ 11.0 11.1 Billing-Ross, Paul; Germain, Arnaud; Ye, Kaixiong; Keinan, Alon; Gu, Zhenglong; Hanson, Maureen R. (January 20, 2016). "Mitochondrial DNA variants correlate with symptoms in myalgic encephalomyelitis/chronic fatigue syndrome". Journal of Translational Medicine. 14 (1): 19. doi:10.1186/s12967-016-0771-6. ISSN 1479-5876.
- ↑ 12.0 12.1 Lande, Asgeir; Fluge, Øystein; Strand, Elin B.; Flåm, Siri T.; Sosa, DaysiD.; Mella, Olav; Egeland, Torstein; Saugstad, OlaD.; Lie, Benedicte A. (March 24, 2020). "Human Leukocyte Antigen alleles associated with Myalgic Encephalomyelitis/Chronic Fatigue Syndrome (ME/CFS)". Scientific Reports. 10 (1): 1–8. doi:10.1038/s41598-020-62157-x. ISSN 2045-2322.
- ↑ 13.0 13.1 Smith, Alicia K.; Fang, Hong; Whistler, Toni; Unger, Elizabeth R.; Rajeevan, Mangalathu S. (2011). "Convergent Genomic Studies Identify Association of GRIK2 and NPAS2 with Chronic Fatigue Syndrome". Neuropsychobiology. 64 (4): 183–194. doi:10.1159/000326692. ISSN 0302-282X. PMC 3701888. PMID 21912186.
- ↑ 14.0 14.1 Dibble, Joshua J; McGrath, Simon J; Ponting, Chris P (August 3, 2020). "Genetic risk factors of ME/CFS: a critical review". Human Molecular Genetics. 29 (R1): R117–R124. doi:10.1093/hmg/ddaa169. ISSN 0964-6906. PMC 7530519. PMID 32744306.
- ↑ 15.0 15.1 15.2 15.3 Hajdarevic, Riad; Lande, Asgeir; Mehlsen, Jesper; Rydland, Anne; Sosa, DaisyD.; Strand, Elin B.; Mella, Olav; Pociot, Flemming; Fluge, Øystein; Lie, Benedicte A.; Viken, Marte K. (May 1, 2022). "Genetic association study in myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS) identifies several potential risk loci". Brain, Behavior, and Immunity. 102: 362–369. doi:10.1016/j.bbi.2022.03.010. ISSN 0889-1591.
- ↑ "TPPP Gene | TPPP Protein | TPPP Antibody". GeneCards. Retrieved March 29, 2022.
- ↑ 17.0 17.1 Goertzel, Benjamin N.; Pennachin, Cassio; de Souza Coelho, Lucio; Gurbaxani, Brian; Maloney, Elizabeth M.; Jones, James F. (April 2006). "Combinations of single nucleotide polymorphisms in neuroendocrine effector and receptor genes predict chronic fatigue syndrome". Pharmacogenomics. 7 (3): 475–483. doi:10.2217/14622416.7.3.475. ISSN 1462-2416. PMID 16610957.
- ↑ Gow, John W.; Hagan, Suzanne; Herzyk, Pawel; Cannon, Celia; Behan, Peter O.; Chaudhuri, Abhijit (June 25, 2009). "A gene signature for post-infectious chronic fatigue syndrome". BMC Medical Genomics. 2 (1): 38. doi:10.1186/1755-8794-2-38. ISSN 1755-8794.
- ↑ Light, A.R.; White, A.T.; Hughen, R.W.; Light, K.C. (2009). "Moderate Exercise Increases Expression for Sensory, Adrenergic, and Immune Genes in Chronic Fatigue Syndrome Patients But Not in Normal Subjects". The Journal of Pain. 10: 1099–1112. PMC 2757484.
- ↑ Albright, Frederick; Light, Kathleen; Light, Alan; Bateman, Lucinda; Cannon-Albright, Lisa A (2011). "Evidence for a heritable predisposition to Chronic Fatigue Syndrome". BMC Neurology. 11 (62). doi:10.1186/1471-2377-11-62.
- ↑ Light, A.R.; Bateman, L.; Jo, D.; Hughen, R. W.; VanHaitsma, T. A.; White, A. T.; Light, K. C. (July 13, 2011). "Gene expression alterations at baseline and following moderate exercise in patients with Chronic Fatigue Syndrome and Fibromyalgia Syndrome". Journal of Internal Medicine. 271 (1): 64–81. doi:10.1111/j.1365-2796.2011.02405.x. ISSN 0954-6820. PMC 3175315. PMID 21615807.
- ↑ Schlauch, Karen A.; Khaiboullina, Svetlana F.; De Meirleir, Kenny L.; Rawat, Shanti; Petereit, J; Rizvanov, Albert A; Blatt, Nataliya; Mijatovic, Tatjana; Kulick, D; Palotás, András; Lombardi, Vincent C. (2016). "Genome-wide association analysis identifies genetic variations in subjects with myalgic encephalomyelitis/chronic fatigue syndrome". Translational Psychiatry. 6 (2): e730. doi:10.1038/tp.2015.208. PMC 4872418.
- ↑ Trivedi, Malav S.; Oltra, Elisa; Sarria, Leonor; Rose, Natasha; Beljanski, Vladimir; Fletcher, Mary Ann; Klimas, Nancy G.; Nathanson, Lubov (2018). "Identification of Myalgic Encephalomyelitis/Chronic Fatigue Syndrome-associated DNA methylation patterns". PloS One. 13 (7): e0201066. doi:10.1371/journal.pone.0201066. ISSN 1932-6203. PMID 30036399.
- ↑ Herrera, Santiago; de Vega, Wilfred C.; Ashbrook, David; Vernon, SuzanneD.; McGowan, Patrick O. (December 5, 2018). "Genome-epigenome interactions associated with Myalgic Encephalomyelitis/Chronic Fatigue Syndrome". Epigenetics: 1–17. doi:10.1080/15592294.2018.1549769. ISSN 1559-2308. PMID 30516085.
- ↑ Nguyen, Chinh Bkrong; Kumar, Surendra; Zucknick, Manuela; Kristensen, Vessela N.; Gjerstad, Johannes; Nilsen, Hilde; Wyller, Vegard Bruun (February 2019). "Associations between clinical symptoms, plasma norepinephrine and deregulated immune gene networks in subgroups of adolescent with Chronic Fatigue Syndrome". Brain, Behavior, and Immunity. 76: 82–96. doi:10.1016/j.bbi.2018.11.008. ISSN 1090-2139. PMID 30419269.
- ↑ Cheema, Amanpreet K.; Sarria, Leonor; Bekheit, Mina; Collado, Fanny; Almenar‐Pérez, Eloy; Martín‐Martínez, Eva; Alegre, Jose; Castro‐Marrero, Jesus; Fletcher, Mary A.; Klimas, Nancy; Oltra, Elisa; Nathanson, Lubov (April 14, 2020). "Unravelling myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS): Gender-specific changes in the microRNA expression profiling in ME/CFS". Journal of Cellular and Molecular Medicine. 00: 1–13. doi:10.1111/jcmm.15260. ISSN 1582-4934. PMID 32291908.
- ↑ Fernandez-Guerra, Paula; Gonzalez-Ebsen, Ana C.; Boonen, Susanne E.; Courraud, Julie; Gregersen, Niels; Mehlsen, Jesper; Palmfeldt, Johan; Olsen, Rikke K.J.; Brinth, Louise Schouborg (July 2021). "Bioenergetic and Proteomic Profiling of Immune Cells in Myalgic Encephalomyelitis/Chronic Fatigue Syndrome Patients: An Exploratory Study". Biomolecules. 11 (7): 961. doi:10.3390/biom11070961. ISSN 2218-273X.
- ↑ Bedford, Felice L.; Greshake Tzovaras, Bastian (March 18, 2021). "Re-analysis of Genetic Risks for Chronic Fatigue Syndrome From 23andMe Data Finds Few Remain". Frontiers in Pediatrics. 9: 590040. doi:10.3389/fped.2021.590040. ISSN 2296-2360. PMC 8012483. PMID 33816394.

